Learning the language of proteins with transformers
May 16, 2023
Authors

Kshitij Vichare
Senior Data Scientist at Fractal, AIML-ADM
Take a trip into the future of bioinformatics with Kshitij Vichare, Senior Data Scientist at Fractal, and discover the impact of transformers on three-dimensional protein models.
Proteins are essential molecules that carry out many biological functions in living organisms, making them crucial targets for biotechnology research. Studying proteins can provide insights into their structures, interactions, and functions. Specialists can thus develop new drugs, therapies, and diagnostic tools by digging deeper into their structure. Biotechnology researchers can apply protein engineering, recombinant DNA technology, and proteomics to manipulate and study proteins. Engineered proteins can boast specific functions or properties, affording exceptional stability, activity, and specificity. A comprehensive understanding of proteins is indispensable in understanding illnesses as various diseases arise from protein dysfunction or misfolding.
Advances in biotech have made it possible to sequence and analyse entire genomes, leading to the discovery of new protein families. Artificial intelligence (AI) is becoming increasingly significant in this research.
Recent advances in artificial intelligence (AI) have created possibilities for exciting applications in protein research. The application of AI in predicting three-dimensional (3D) protein structure and function from amino acid sequences and designing proteins with specific functions through protein engineering is of particular interest. AI-based analysis of protein data sets also enables the identification of novel biomarkers for disease diagnosis and monitoring. These developments have the potential to uncover new insights into disease mechanisms and lead to the development of more effective treatments that would be difficult — if not impossible — without the assistance of AI.
Challenges Facing the Industry
Although technology is helping move protein-specific research forward, the industry still faces several challenges:
Limited representation of protein structure
Proteins are complex molecules that fold into unique three-dimensional structures essential for their functions. However, traditional methods for representing protein structures, such as using the amino acid sequences that make up the protein, need to provide the necessary information for modeling proteins to understand how proteins work entirely. Researchers are developing new methods to represent protein structures, such as using three-dimensional models, molecular dynamics simulations, or hybrid approaches that combine sequence and structural information. These methods are expected to improve the accuracy of protein models in predicting protein properties and functions.
Lack of context-dependent features
Protein structure and function depend highly on the interactions between amino acids and structural motifs. However, traditional representations of protein structure often need to capture these crucial details making it difficult for protein models to make accurate predictions. To push boundaries in protein research, leading-edge approaches are being explored for contextualizing models. These include attention mechanisms that focus on segments of the proteins and graph neural networks to detect correlations among amino acids and motifs.
Difficulty in handling large protein structures
Proteins can be very large and complex molecules, making it difficult for protein models to process them efficiently. This can lead to problems with training and optimizing the relevant models. Innovative approaches are being explored to handle large protein structures, such as using hierarchical approaches that break down the protein into smaller substructures or using specialized hardware such as Graphical Processing Units (GPUs) or Tensor Processing Units (TPUs) that can accelerate the computation of protein models. These methods are expected to improve the scalability and efficiency of protein models, allowing them to process larger and more complex protein structures.
Recently, researchers have begun using transformers to improve the accuracy of protein models. The results have been exciting and address many of the abovementioned issues.
The Power of Transformers
The transformer is an innovative deep learning model that employs powerful attention mechanisms to process sequential input data. The central innovation in the neural architecture of transformers is the self-attention mechanism, which allows the entire sequence to be processed simultaneously rather than sequentially, as other models do. This allows for long-range dependencies in the sequence and has been used to achieve state-of- the-art accuracy in natural language processing (NLP) tasks such as text classification, sentiment analysis, and language translation.
Transformers are like language models that extract concepts from sequences and build models around them, while protein language models use evolutionary sequences to build models based on interactions between residues.
Transformers use multi-headed attention to transform each token sequence into key-value pairs and then map each key to the value of other tokens in the sequence, generating context vectors for each specific token. Applying this to protein structures allows them to process evolutionary sequences and consider their context to find interactions between amino acid residues. The ability to do this results in more accurate protein structure predictions.
Transformer architectures enable sophisticated parallel processing, allowing entire sequences to be processed simultaneously and facilitating the capture of intricate relationships between elements within the input. This makes it possible to train the model on vast amounts of data, improving its accuracy.
Additionally, pre-training can be done by masking random tokens in the sequence and training the model to predict them, allowing for non-annotated text data to retrain the model. This was impossible with previous language models, thereby making transformers a powerful tool in natural language processing and protein language modeling.
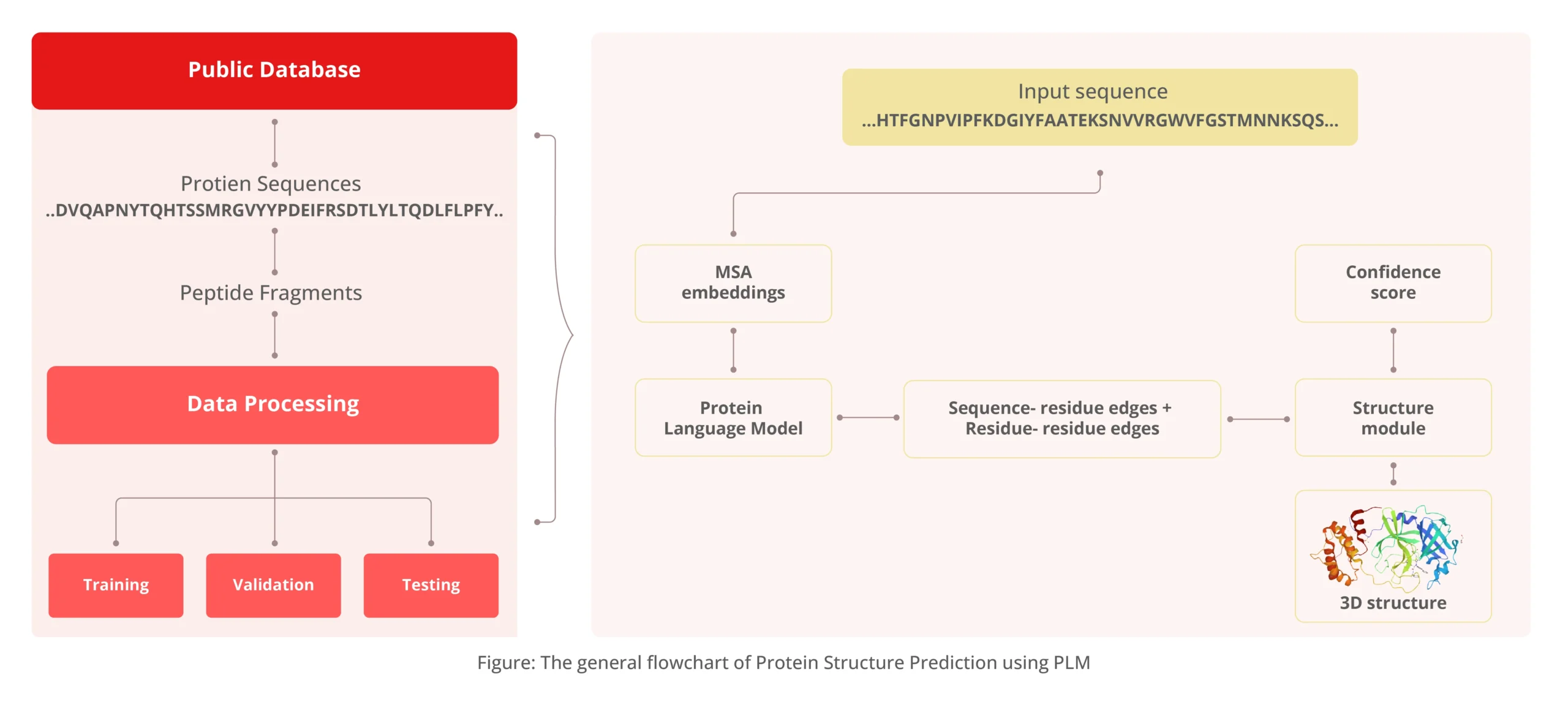
The figure above describes an ML pipeline to predict protein structure from amino acid sequences. The transformer-based model considers the evolutionary relationships between protein sequences to generate a structure that accurately represents the protein.
Creating next-level 3D protein models
Representing protein structures as three-dimensional graphs captures the spatial relationships between amino acids, providing additional information beyond their linear sequence. Two approaches currently being used to create these 3D models are graph attention networks (GAT) and graph convolutional networks (GCN).
GATs encode the graph structure of the protein using multi-headed attention, allowing the model to scan the entire graph simultaneously and focus on the most essential amino acid interactions. Meanwhile, GCNs update the node embeddings with local neighborhood aggregation to encode the graph structure of the protein.
Both approaches transform the graph information into the three-dimensional model by adding a layer that processes the graph structure of the protein and then combines it with the output of the transformer layer that processes the amino acid sequence. This technique is novel because it combines the power of transformers with the ability of graph structures to capture spatial relationships between amino acids.
GATs have performed better than GCNs because of their ability to capture more complex relationships between amino acids and long-term dependencies — including non-covalent bonds. However, both approaches have shown promising results in predicting protein structure and function. Combining GATs and GCNs with transformers promises new insights into the relationship between protein structure and function.
Fulfilling the data needs of transformers
In bioinformatics, clustering algorithms are crucial in analysing and interpreting complex biological data, including proteins. Clustering of proteins with unknown characteristics has a useful application involving grouping proteins based on similarities in their properties or behavior. By doing so, researchers can identify patterns and similarities between different proteins, which can provide insights into their function, structure, and interactions with other molecules.
Clustering and identifying patterns in proteins with unknown characteristics can help develop protein modeling with transformers. This approach requires an extensive pre-training dataset to create protein structures using transformers. However, obtaining and annotating a sizeable pre-training dataset can be challenging and resource intensive. By leveraging clustering and pattern recognition algorithms, researchers can identify similarities and patterns between proteins with known functions and those with unknown characteristics. This information can augment pre-training datasets, enabling the creation of more accurate and precise models.
In addition, once a pre-trained model has been created, transfer learning can be utilized to fine-tune the model for specific tasks. Transfer learning involves taking the knowledge and patterns learned from the pre-trained model and applying it to a new task, such as protein classification or prediction of protein-ligand binding affinity.
By leveraging clustering algorithms and transfer learning, researchers can develop more accurate and efficient models for protein analysis without requiring extensive, labeled datasets for each specific task. This approach can lead to more effective drug discovery, protein engineering, personalized medicine, and a better understanding of the fundamental biological processes underlying human health and disease.
The Future of Bioinformatics
Transformers have emerged as a powerful tool in developing state-of-the-art protein models, including the AlphaFold2 model, which achieved ground-breaking results in the 2020 protein folding prediction competition. These models offer exciting possibilities for advancing our understanding of proteins and their functions in bioinformatics.
Despite their success, there is still ongoing research in developing these models. One area of interest is the ability of transformers to process multi-modal data structures or information. This would involve generating a latent space vector from the entire structure of a protein, which could then be used to predict what the structure would look like given a novel amino acid sequence. Researchers could develop even more accurate and robust protein models by combining information from different modalities.
Fractal is exploring different approaches to multi-modal learning through transformers focusing on generating latent space vectors. This has already been successful in text-to-image generation models, and Fractal is interested in applying this concept to protein structure prediction. Transformers have significantly contributed to bioinformatics and protein language model development. However, ongoing research in the field, such as using multi-modal data structures, can further advance our understanding of proteins and their functions, ultimately leading to more effective drug discovery and personalized medicine.
Recognition and achievements





